Keywords: Robust Growth Curve Modeling · Bayesian Estimation · Structural Equation Modeling · JAGS
Latent growth curve models (LGCM) are widely used in longitudinal studies, and LGCM performs well in the identification of intraindividual changes and investigation of interindividual differences in intraindividual changes (McArdle & Nesselroade, 2014). LGCM can estimate linear and nonlinear growth trajectories flexibly or freely estimate the shape of growth trajectory by observed data. Researchers may employ either the maximum likelihood estimation method or the Bayesian method to model LGSM. The Bayesian methods have advantages on handling difficulties in longitudinal data such as unequally spaced measurements, nonlinear trajectories, non-normally distributed data, and small sample sizes (Curran, Obeidat, & Losardo, 2010).
Influential outliers and non-normally distributed data can lead unreliable estimates and inferences. Conventional methods such as deleting outliers may result in underestimated standard errors (Lange, Little, & Taylor, 1989). Robust statistical modeling methods have been developed to handle the violation of the normality assumption. For example, the \(t\)-distribution is more robust to outliers, and using \(t\)-distribution to model errors is one of the robust modeling strategies (Lange et al., 1989). Robust modeling using \(t\)-distributions is easy to understand and applied in both maximum likelihood and Bayesian methods (Lange et al., 1989; Zhang, 2016; Zhang, Lai, Lu, & Tong, 2013). The degree of freedom of \(t\)-distributions can be estimated or predetermined, and a large degree of freedom means the \(t\)-distribution approaches a normal distribution (Zhang et al., 2013). Based on simulation studies, the robust method using the \(t\)-distributions for the error term demonstrates good performance for heavy-tailed data in growth curve models, and it efficiently estimates the standard error (Zhang, 2016; Zhang et al., 2013).
This tutorial aims to present how to implement robust Bayesian growth curve models using R and the JAGS programs. To begin, it provides a brief introduction to LGCM, including the latent basis growth curve models (LBGM), the linear growth curve models, and quadratic growth curve models. Then it introduces commonly used priors and convergence diagnostic methods. Finally, a real data set is used to demonstrate how to implement robust LGCM, and how to interpret the estimated parameters.
A LGCM with one variable \(Y\) can be written as:
\begin {equation} \mathbf {Y}_i = \boldsymbol {\tau } + \mathbf {\Lambda } \boldsymbol {b}_i + \boldsymbol {\epsilon }_i \end {equation}
\begin {equation} \boldsymbol {b}_i = \boldsymbol {\beta } + \boldsymbol {u}_i \end {equation} \(\mathbf {Y}_i\) is a \(T \times 1\) vector in which \(T\) is the total number of measurement occasions, and \(\mathbf {\Lambda }\) is a \(T \times q\) factor loading matrix, and it decides the shape of the growth trajectory. The \(\boldsymbol {\epsilon }_i\) is assumed to follow a \(q\)-variate normal distribution \(\boldsymbol {\epsilon }_i \sim \text {MN}(0,\mathbf {\Phi })\). \(\boldsymbol {b}_i\) is an \(q \times 1\) vector and it represents the latent variables used to describe the change. \(\boldsymbol {\beta }\) is a \(q \times 1\) vector that represents the fixed effect (the means of \(\boldsymbol {b}_i\)) and \(\boldsymbol {e}_i\) is the individual deviation from the fixed effect \(\boldsymbol {\beta }\). \(\boldsymbol {u}_i\) follows a multivariate normal distribution with \(q\) dimensions as \(\boldsymbol {u}_i \sim \text {MN}(0,\mathbf {\Psi })\).
LBGM is a special case of the general LGCM. It assumes the error variance is the same for all measurements (homogeneity) by simplifying \(\mathbf {\Phi }=\mathbf {I} \sigma _e^2\). And it also assumes measurement errors are uncorrelated. The parameters in LBGM are:
\begin {align*} \mathbf {\Lambda } = \begin {pmatrix} 1 & 0\\ 1 & 1\\ 1 & \lambda _1\\ \vdots & \vdots \\ 1 & \lambda _{T-2} \end {pmatrix} & & \boldsymbol {b}_i = \begin {pmatrix} b_{iL}\\ b_{iS} \end {pmatrix}, \\ \boldsymbol {\beta } = \begin {pmatrix} \beta _{L}\\ \beta _{S} \end {pmatrix} & & \mathbf {\Psi }= \begin {pmatrix} \sigma ^2_L & \sigma _{LS}\\ \sigma _{LS} & \sigma ^2_S \end {pmatrix}. \end {align*}
LBGM contains two latent variables: \(b_L\) and \(b_S\). \(b_L\) represents the intercept and \(b_S\) represents the growth slope. The specification of factor loadings of \(b_S\) determines the shape of the growth curve. Here, the first and second-factor loadings on \(b_S\) are fixed at 0 and 1 for identification purposes, while other factor loadings are freely estimated. This assumption implies that the growth unit is the difference between the first two measurements. Another common practice is to fix the first and last factor loadings at 0 and 1, respectively, with the unit representing the difference between the first and last measurements. \(\beta _L\) and \(\beta _S\) represent the average intercept and slope across all individuals, respectively. \(\sigma ^2_L\) and \(\sigma ^2_S\) represent variances, reflecting the individual differences in intercept and slope. \(\sigma _{LS}\) represents the covariance between the intercept and slope.
The specification of \(\mathbf {\Lambda }\) decides the shape of growth. When the factor loadings of \(b_S\) are equally spaced, it becomes a linear growth curve model. A linear growth curve model assumes a linear change pattern and the slope \(b_S\) represents a linear slope. The factor loading matrix is:
\[\mathbf {\Lambda } = \begin {pmatrix} 1 & 0\\ 1 & 1\\ 1 & 2\\ \vdots & \vdots \\ 1 & T-1 \end {pmatrix}.\]
The quadratic growth curve model estimates a nonlinear change by including the quadratic slope \(b_{iQ}\), and the model can be presented as:
\begin {align*} \mathbf {\Lambda } = \begin {pmatrix} 1 & 0 & 0 \\ 1 & 1 & 1 \\ 1 & 2 & 2^2 \\ \vdots & \vdots & \vdots \\ 1 & T-1 & {(T-1)}^2 \end {pmatrix} & & \boldsymbol {b}_i = \begin {pmatrix} b_{iL}\\ b_{iS}\\ b_{iQ} \end {pmatrix}, \\ \boldsymbol {\beta } = \begin {pmatrix} b_{iL}\\ b_{iS}\\ b_{iQ} \end {pmatrix} & & \mathbf {\Psi }= \begin {pmatrix} \sigma ^2_L & \sigma _{LS} & \sigma _{LQ}\\ \sigma _{LS} & \sigma ^2_S & \sigma _{SQ}\\ \sigma _{LQ} & \sigma _{SQ} & \sigma _Q^2 \end {pmatrix} . \end {align*}
The general LGCM assumes \(\boldsymbol {\epsilon }_i\) follow a multivariate normal distribution (\(\boldsymbol {\epsilon }_i \sim MN(0,\mathbf {\Phi })\)), while robust growth curve models use other distributions to for \(\boldsymbol {\epsilon }_i\). Zhang (2016) presented and summarized how to use Student’s \(t\), exponential power, and the skew normal distributions to build robust LGCM.
The robust growth curve models can be specified by modeling \(\boldsymbol {\epsilon }_i\) by a student’s \(t\)-distribution: \(\boldsymbol {\epsilon }_i \sim MT_T(0,\Phi ,k)\), where \(k\) is the degrees of freedom. The robust LGCM with a Students’ \(t\)-distribution performs better than the traditional growth curve model with a multivariate normal distribution when dealing with heavy-tailed data and outliers (Zhang, 2016; Zhang et al., 2013).
The multivariate \(t\)-distribution approaches the multivariate normal distribution when \(k\) increases. In the robust Bayesian methods, \(k\) can be specified as an unknown parameter, and a prior is needed to estimate \(k\). Alternatively, it can be fixed and some researchers suggested \(k=5\) (Zhang et al., 2013).
In JAGS, \(t\)-distribution can be specified using the function dt(), and this function will be explained in the following section with an example.
The exponential power distribution can model error term \(e_{it}\) with smaller kurtosis than normal distributions, and we employ the same form of density function and parameters as Zhang (2016) in this tutorial. The density of exponential power distribution is as follows:
\[p_{ep}(x)=\omega (\gamma )\sigma ^{-1}exp\left [-c(\gamma )\big |\cfrac {x-\mu }{\sigma } \big |^{2/(1+\gamma )}\right ]\] where \[\omega (\gamma )=\cfrac {\left ( \Gamma \left [3(1+\gamma )/2\right ]\right )^{1/2} }{(1+\gamma )\left (\Gamma \left [(1+\gamma )/2\right ]\right )^{3/2}}\] and \[c(\gamma )=\left ( \cfrac {\Gamma [3(1+\gamma )/2]}{\Gamma [(1+\gamma )/2]}\right )^{1/(1+\gamma )}.\] Here \(\mu \) and \(\sigma \) are location and scale parameters, respectively, and \(\gamma \) is a shape parameter that can be estimated.
Both the \(t\)-distribution and the exponential power distribution are symmetric, while the skew normal distribution offers an option to model asymmetric errors. The density function of a skew normal distribution is as follows:
\[p_{sn}(x)=\cfrac {2}{\omega }\phi \left (\cfrac {x-\mu }{\omega }\right )\Phi \left (\alpha \cfrac {x- \mu }{\omega }\right )\] where \(\mu \) is a location parameter, \(\omega \) is a scale parameter, and \(\alpha \) is a shape parameter which can be estimated.
The following part introduces how to build and interpret the robust LGCM in JAGS using a real data set, assuming homogeneity across time points.
Priors of LGCM are usually specified as: \(\boldsymbol {\beta } \sim N(\mu _0,\sigma _0^2)\), \(\mathbf {\Phi } \sim W(V,m)\), \({\sigma _e^2} \sim IG(\alpha ,\beta )\) (assuming \(\mathbf {\Phi }=\mathbf {I}_{T \times T}\sigma _e^2\)). For the robust growth curve model with the \(t\)-distribution, the degrees of freedom \(k\) is another unknown parameter, and an uninformative prior is applied to \(k\) as follows: \(k \sim U(1,500)\). In the case of the exponential power distribution which involves an additional shape parameter \(\gamma \), an uninformative prior is assigned to it as follows: \(\gamma \sim U(-1,1)\). Similarly, for the shape parameter \(\alpha \) in the skew normal distribution, the prior is specified as \(\alpha \sim U(-5,5)\).
To check convergence, trace plots are visually inspected. If trace plots indicate non-convergence, then more iterations and longer burn-in periods are needed. The length of the chain should be extended until trace plots of all parameters demonstrate visual convergence.
In addition to visual inspection, various convergence diagnostic tools are available in R, including the Geweke test (Geweke, 1992), the Heidelberger and Welch test (Heidelberger & Welch, 1983), Gelman and Rubin test (Gelman & Rubin, 1992), and the Raftery and Lewis diagnostic (Raftery, Lewis, et al., 1992). In this tutorial, the Geweke diagnostic is used, which compares the mean difference between two parts of chains, typically the first and last parts. It employs a \(z\) test to compare the means of two parts, and if the \(z\) test statistic rejects the null hypothesis, it indicates a significant difference.
The adjacent iterations of the Markov chain may exhibit high dependence, and serious autocorrelation can indicate problems in model estimation such as a problem with the sampling algorithm. The autocorrelation problem can be identified by visual inspections. If visual inspection shows high autocorrelation, increasing the number of iterations or implementing thinning techniques can be beneficial. Additionally, it is important to ensure that the posterior distribution makes substantive sense, taking into account factors such as the parameter’s range and standard deviation. For instance, it would be unreasonable if the posterior standard deviation exceeds the parameter’s scale.
This section includes R code and JAGS commands for constructing robust growth curve models. The \(t\)-distribution is offered by JAGS and can be directly implemented. In the following parts, \(t\)-distribution is utilized to model and compare LBGC, linear and quadratic LGCM. To illustrate different robust methods, we specify linear LGC models using the \(t\), exponential power and skew-normal distributions.
The data used in this tutorial were obtained from the Early Childhood Longitudinal Study, Kindergarten Class of 2010-11 (ECLS-K:2011), a national longitudinal program conducted by the National Center for Education Statistics. ECLS-K:2011 collected information about children’s development during their elementary school years. For this tutorial, a random subset of data consisting of \(N=200\) samples was selected from ECLS-K:2011. This subset includes math scores measured at four different occasions. Math ability assessments were conducted annually, spanning from the second grade to the fifth grade. Detailed information about ECLS-K:2011 can be found in the manual provided by Tourangeau et al. (2015).
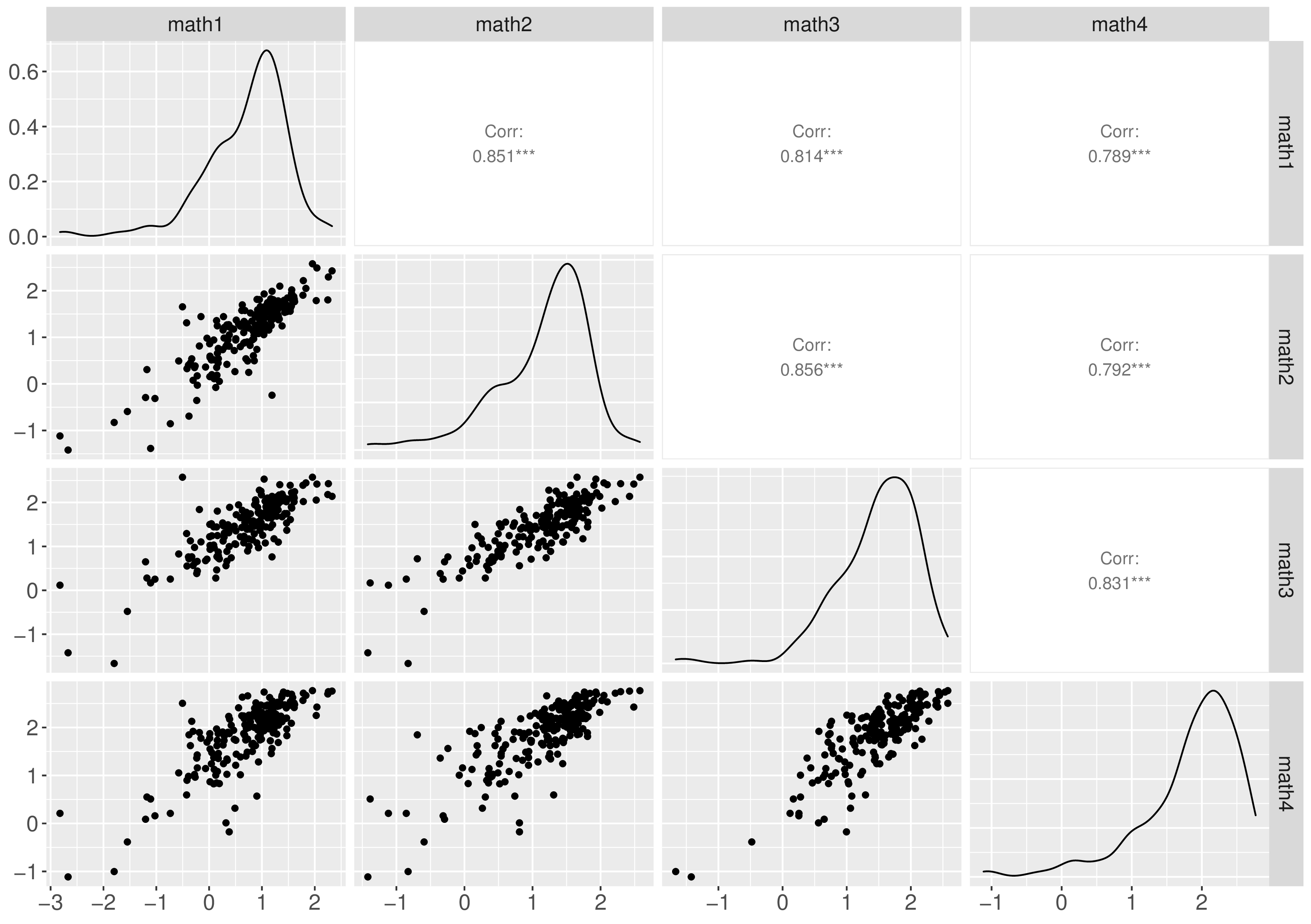
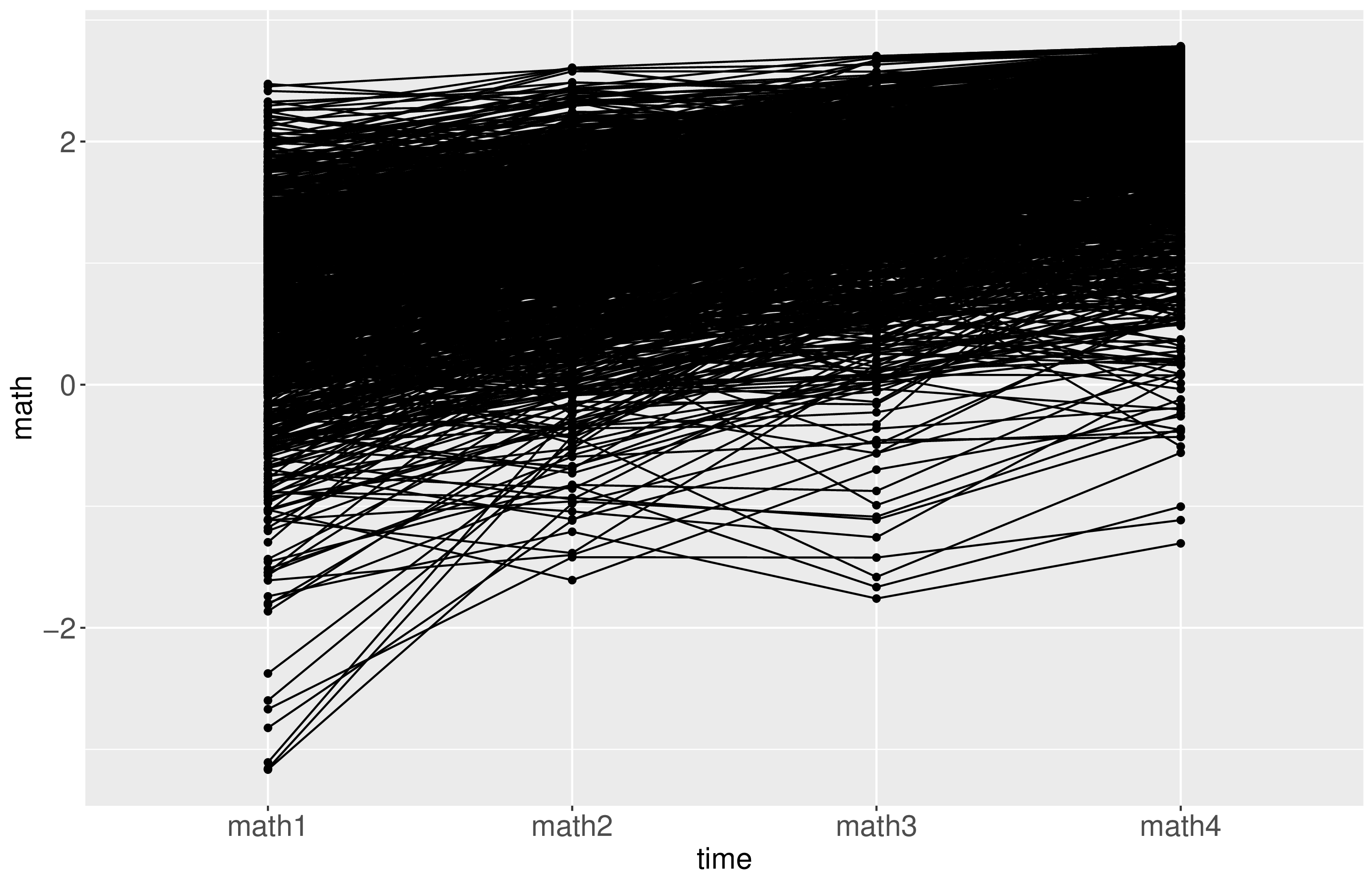
Descriptive analysis revealed that the distributions of the observed math scores were skewed and exhibited heavier tails than normal distributions, as depicted in Figure 1. Additionally, increasing trends in math scores were observed, and the growth pattern of each individual is illustrated in Figure 2.

The LBGM model is specified using the JAGS notations as:
# models model1 <- "model{ # Specify the likelihood for (i in 1:nsubj) { for (j in 1:ntime) { # t error y[i, j] ˜ dt(mu[i, j], tauy, df) # normal # y[i, j] ˜ dnorm(mu[i, j],tauy) } } for (i in 1:nsubj){ mu[i,1] <- b[i,1] mu[i,2] <- b[i,1]+b[i,2] mu[i,3] <- b[i,1]+A3*b[i,2] mu[i,4] <- b[i,1]+A4*b[i,2] b[i,1:2] ˜ dmnorm(mub[1:2], taub[1:2,1:2]) } # Specify the growth trajectory A3˜dnorm(0,1.0E-6) A4˜dnorm(0,1.0E-6) # specify priors mub[1]˜dnorm(0,1.0E-6) mub[2]˜dnorm(0,1.0E-6) taub[1:2, 1:2] ˜ dwish(Omega[1:2, 1:2], 2) sigma2b[1:2, 1:2] <- inverse(taub[1:2, 1:2]) tauy ˜ dgamma(0.001,0.001) sigma2y <- 1 / tauy df ˜ dunif(1,500) Omega[1,1] <- 1 Omega[2,2] <- 1 Omega[1,2] <- Omega[2,1] Omega[2,1] <- 0 } "
An R object model1 is constructed using the model block. In the case of a four-wave of data organized with 200 rows (\(N=200\)) and 4 columns (\(T=4\)), we use for loops to specify the likelihood for all participants across the four measurement occasions.
This likelihood reflects the use of a robust Bayesian method. Specifically, y[i,j] is modeled using univariate \(t\)-distributions, which are defined by dt() with parameters for means mu[i, j], precision tauy, and degrees of freedom df. If the data were modeled using a multivariate normal distribution, i.e., y[i,j] \(\sim \) dnorm(mu[i,j], tauy), the model would represent a traditional latent growth curve model with normal assumptions.
The next part of the model involves specifying the prior distributions. \(\boldsymbol {\beta }\) is assumed to follow a bivariate normally distribution with \(\boldsymbol {\beta } \sim MN((0,0)^T,1000\boldsymbol {I_2})\), and the covariance of \(\boldsymbol {\epsilon }_i\) follows an inverse Wishart distribution (Zhang, 2021). The error term \(\boldsymbol {\epsilon }_i\) is assumed to follow a \(t\)-distribution with an estimated \(k\) (\(\boldsymbol {\epsilon }_i \sim MT_T(0,\mathbf {\Phi },k)\)). Here, a uniform distribution \(Unif(1,500)\) is used as the prior of \(k\).
The latent variables and means are specified based on the hypothesized growth curve and priors. The parameter b[i,1] represents the latent intercept of LGCM, and the latent slope is b[i,2]. A3 and A4 are factor loadings of math scores at the third and fourth measurement occasions, which control the shape of changes.
If A3 is set to 2 and A4 is set to 3, then the model becomes a linear growth curve model. Quadratic LGCM involves three latent variables, within b[i,3] representing the quadratic shape. The coefficients A5 and A6 are fixed at 4 and 9, respectively.
Detailed JAGS models for both the linear and quadratic growth curve models can be found in the appendix.
JAGS does not offer exponential power distribution or the skew-normal distribution by default. However, the likelihood can be specified indirectly using the Bernoulli or the Poisson distributions (Ntzoufras, 2011).
One approach, known as the “zero trick,” utilizes the Poisson distribution. A matrix with the same dimensions of the data is created, with all elements set to zero. The likelihood is reflected in the mean of the Poisson distribution. Assuming observation \(y_i\) follows a new distribution and the log-likelihood is \(l_i=logf(y_i|\theta )\). The model likelihood can be expressed as:
\[f(y|\theta )=\prod _{i=1}^n\cfrac {e^{-(-l_i+c)}(-l_i+C)^0}{0!}=\prod _{i=1}^2f_P(0;-l_i+C).\] In this expression, the mean of the Poisson distribution is a constant (\(C\)) minus the log-likelihood (\(C-l_i\)) and \(C\) is chosen to ensure the mean of the Poisson distribution is always positive.
The one trick sets all observations to one and uses the parameter of the Bernoulli distribution to specify likelihood.
In this paper, the zero tricks were used to specify exponential power and skew-normal distributions, assuming a linear change trajectory.
The model code is provided in the appedix. In the code, the log gamma function is specified using command loggam(), and dpois() is used to sample from the Poisson distribution.
The location parameter of the skew normal distribution is reparameterized as \[\mu =\omega \frac {\alpha }{\sqrt {(1+\alpha ^2)^2}} \sqrt {(2/\pi )}\] to ensure that the mean of the error is zero. In the code, the standard normal cumulative density function is specified by phi() and the log density function of the normal distribution is specified by logdensity.norm().
After configuring the models, we can proceed by organizing the data in a list, specifying initial values, and running the JAGS model.
The data is organized in a wide format and stored in a list called datalist, which includes the number of participants (nsubj) and the number of measurements (ntime). In this setup, we use two chains (nChains = 2), each with a length of 20,000 iterations (nIter = 20000), and a burn-in period of 10,000 iterations (burnInSteps = 10000). Monitored parameters encompass the means and variances of intercepts and slopes, and the shape parameters such as the degrees of freedom. These parameters’ posterior draws will be saved.
# create data set for ' texttt{JAGS} model nsubj = nrow(data) ntime = ncol(data) datalist = list(nsubj=nsubj,ntime=ntime,y=data) # set parameters, adaption, and MCMC chains parameters = c("mub","sigma2b","sigma2y","df","A3","A4") adaptSteps =5000 # Adaptive period burnInSteps = 10000 # Burn-in period nChains = 2 # The number of chains nIter =20000 # The number of kept iterations
Two chains are used in this tutorial, and two sets of initial values are specified:
# specify initial values inits <- list(list(mub=c(0.7,0.4), taub=structure(.Data=c(1,0,0,10), .Dim=c(2,2)), tauy=10,df=3), list(mub=c(0.8,0.5), taub=structure(.Data=c(2,0,0,8), .Dim=c(2,2)), tauy=15,df=5))
The package runjags is used in this tutorial and the function run.jags() is used to read, compile, and run the model, and the model results are saved for later analysis.
# run JAGS model set.seed(1234) out <- run.jags(model=model, monitor=parameters, data=datalist, n.chains=2, inits=inits, method="simple", adapt=adaptSteps, burnin = burnInSteps, sample=nIter, keep.jags.files=TRUE, tempdir=TRUE)
For convergence checking, we examine both trace plots and Geweke’s test. A visual inspection of the trace plots reveals that all parameters have converged after the adaptation and burn-in period. Figure 3 displays the plots of the latent intercept and slope in the LBGM.
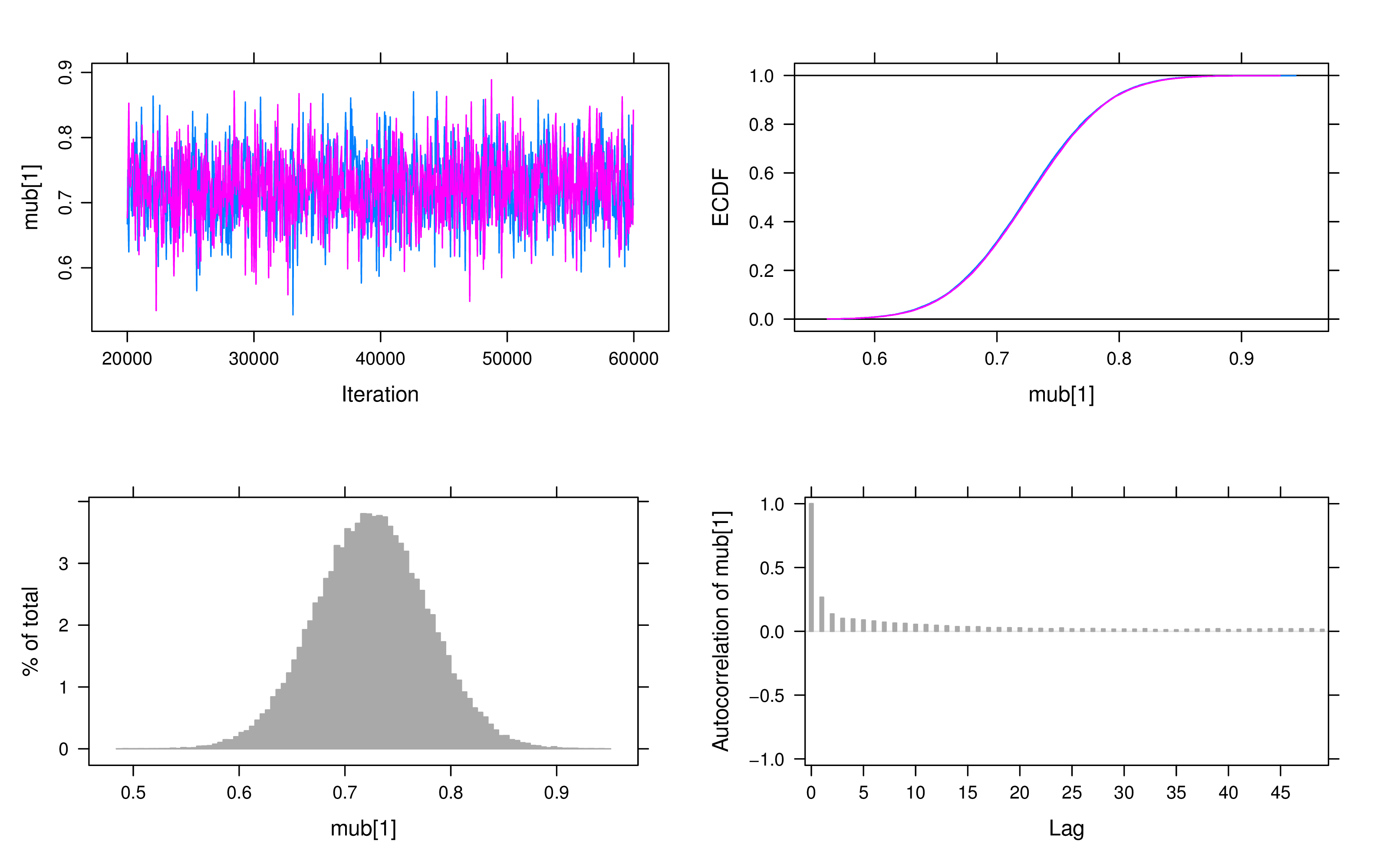
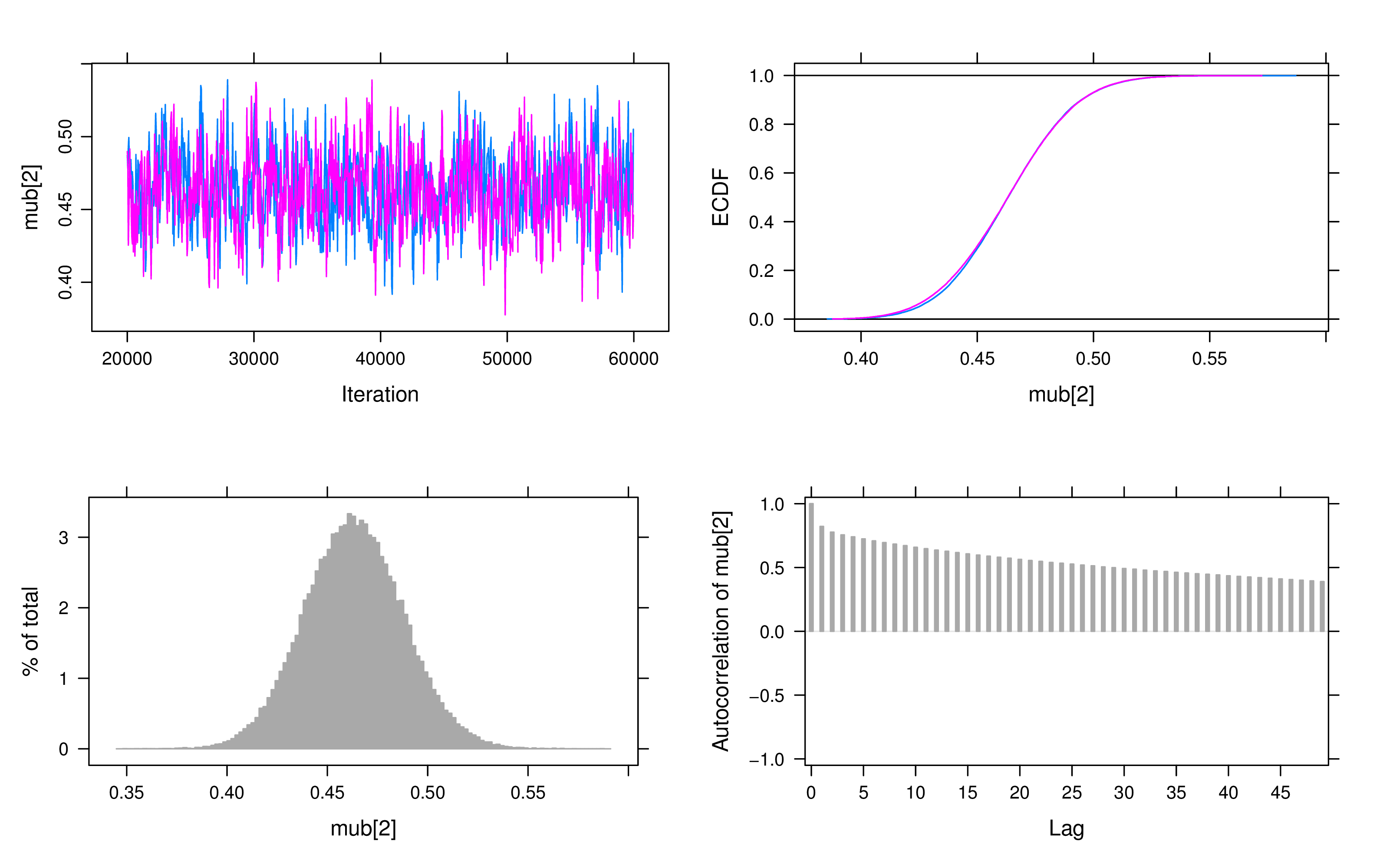
If Geweke’s test values exceeded 2, we doubled the number of iterations and reran the model. In this particular example, we found no clear evidence of non-convergence, however, some models exhibited autocorrelation issues in the slope, as shown in the autocorrelation plots. To address this, longer iterations or thinning techniques may be employed.
Additionally, posteriors make practical sense by checking the shape and range in the posteriors plots. For example, the range of possible values for math ability is from -4.0 to 4.0, and the posterior mean of the intercept was close to the mean of the observed math score in the second grade.
# Geweke diagnostic geweke.diag(out$mcmc) # Trace plots and autocorrelation plots plot(out)
Results of LBGM, the linear and quadratic LGC models are summarized in Table 1. To compare these models, we used the deviance information criterion (DIC). When the \(t\) distribution was employed, the quadratic LGCM exhibited the lowest DIC.
| LBGM | Linear LGCM | Quadratic LGCM
| |||||||
| Mean | L | U | Mean | L | U | Mean | L | U | |
| \(b_{iL}\) | 0.73 | 0.62 | 0.83 | 0.76 | 0.65 | 0.85 | 0.74 | 0.64 | 0.85 |
| \(b_{iS}\) | 0.46 | 0.42 | 0.51 | 0.37 | 0.34 | 0.39 | 0.42 | 0.35 | 0.48 |
| \(b_{iQ}\) | -0.02 | -0.04 | 0.01 | ||||||
| \(\sigma ^2_{L}\) | 0.49 | 0.38 | 0.59 | 0.48 | 0.38 | 0.59 | 0.54 | 0.42 | 0.66 |
| \(\sigma _{LS}\) | -0.05 | -0.07 | -0.02 | -0.04 | -0.06 | -0.02 | -0.11 | -0.18 | -0.05 |
| \(\sigma _{LQ}\) | 0.02 | 0.00 | 0.04 | ||||||
| \(\sigma _{LS}\) | -0.05 | -0.07 | -0.02 | -0.04 | -0.06 | -0.02 | -0.11 | -0.18 | -0.05 |
| \(\sigma ^2_{S}\) | 0.03 | 0.02 | 0.03 | 0.02 | 0.02 | 0.03 | 0.09 | 0.05 | 0.13 |
| \(\sigma _{SQ}\) | -0.02 | -0.04 | -0.01 | ||||||
| \(\sigma _{LQ}\) | 0.02 | 0.00 | 0.04 | ||||||
| \(\sigma _{SQ}\) | -0.02 | -0.04 | -0.01 | ||||||
| \(\sigma ^2_{Q}\) | 0.01 | 0.01 | 0.02 | ||||||
| \(\sigma ^2_{e}\) | 0.03 | 0.02 | 0.04 | 0.03 | 0.02 | 0.04 | 0.03 | 0.02 | 0.04 |
| \(k\) | 3.44 | 2.30 | 4.76 | 3.53 | 2.31 | 4.94 | 3.58 | 2.04 | 5.42 |
| A3 | 1.64 | 1.51 | 1.77 | 2.00 | 2.00 | ||||
| A4 | 2.44 | 2.26 | 2.64 | 3.00 | 3.00 | ||||
| A5 | 4.00 | ||||||||
| A6 | 9.00 | ||||||||
| DIC | 370.79 | 374.78 | 283.36 | ||||||
Note. \(k\) represents the degrees of freedom. L: 2.5% HPD; U: 97.5% HPD. | |||||||||
The estimated means of the intercept \(b_{iL}\) from the three models were close. The estimated factor loadings in LBGM were 1.64 and 2.44 in LBGM, which suggests the estimated growth shape was different from a linear trend.
The estimated degrees of freedom were smaller than 5 in the three models. This aligns with the observation that the observed data had heavier tails than the normal distribution, as shown in Figure 1(Tong & Zhang, 2017). Therefore, the estimated degrees of freedom (\(k\)) are consistent with descriptive statistics, affirming that the robust growth curve models are suitable for handling this dataset.
When dealing with models that use exponential power and skew normal distributions, it’s important to interpret the DIC (deviance information criterion) values from JAGS with caution. In these models, the DIC is calculated separately based on likelihood and posteriors. The deviance, denoted as \(D(\theta ;y)\), is defined as \(-2\log (p(x|\theta ))\). The effective model parameters is defined as \(p_D = \bar {D} - \hat {D}\), and the DIC is calculated as \(DIC = \bar {D} + p_D\). The model using the skew normal distribution exhibited the lowest DIC value, making it the preferred choice over the \(t\) and exponential power distributions.
| Mean | 2.5% HPD | 97.5% HPD | DIC | |
| t-distribution | ||||
| \(b_{iL}\) | 0.76 | 0.65 | 0.85 | |
| \(b_{iS}\) | 0.37 | 0.34 | 0.39 | |
| \(\sigma _{L}^{2}\) | 0.48 | 0.38 | 0.59 | |
| \(\sigma _{LS}\) | -0.04 | -0.06 | -0.02 | 374.78 |
| \(\sigma _{LS}\) | -0.04 | -0.06 | -0.02 | |
| \(\sigma _{S}^{2}\) | 0.02 | 0.02 | 0.03 | |
| \(\sigma _{e}^{2}\) | 0.03 | 0.02 | 0.04 | |
| \(k\) | 3.53 | 2.31 | 4.94 | |
| Exponential power distribution | ||||
| \(b_{iL}\) | 0.76 | 0.66 | 0.86 | |
| \(b_{iS}\) | 0.37 | 0.34 | 0.4 | |
| \(\sigma _{L}^{2}\) | 0.49 | 0.39 | 0.6 | |
| \(\sigma _{LS}\) | -0.04 | -0.06 | -0.02 | 392.10 |
| \(\sigma _{LS}\) | -0.04 | -0.06 | -0.02 | |
| \(\sigma _{S}^{2}\) | 0.02 | 0.02 | 0.03 | |
| \(\sigma _{e}^{2}\) | 0.07 | 0.06 | 0.08 | |
| \(\gamma \) | 0.91 | 0.76 | 1 | |
| Skew normal distribution | ||||
| \(b_{iL}\) | 0.74 | 0.64 | 0.83 | |
| \(b_{iS}\) | 0.37 | 0.35 | 0.4 | |
| \(\sigma _{L}^{2}\) | 0.46 | 0.36 | 0.56 | |
| \(\sigma _{LS}\) | -0.04 | -0.06 | -0.02 | 360.41 |
| \(\sigma _{LS}\) | -0.04 | -0.06 | -0.02 | |
| \(\sigma _{S}^{2}\) | 0.02 | 0.02 | 0.03 | |
| \(\sigma _{e}^{2}\) | 0.18 | 0.15 | 0.21 | |
| \(\alpha \) | -4.17 | -5 | -3.01 | |
The posterior means of most parameters were almost the same for linear models using \(t\), exponential power, and skew-normal distributions, see Table 2. For the linear LGCM with the exponential power error, the estimated shape parameter \(\gamma \) was 0.91, which suggested a fatter tail of the errors than the normal distribution. The estimated \(\alpha \) in the model using the skew-normal distribution was -4.17 which indicates the distribution was left-skewed.
LGCM is widely used in longitudinal studies, and the Bayesian approach can be applied to handle complex conditions. Bayesian approaches can handle the conditions that data are not normally distributed or the sample size is small. The robust Bayesian method offers an operable solution for data with heavy tails or outliers.
This tutorial introduces how to implement robust LGCM with three distributions in JAGS and R in steps. It also covers the model diagnostics and comparison, and interpretations of posterior estimations. This tutorial offers some guidelines for researchers who are interested in robust Bayesian growth curve models.
Curran, P. J., Obeidat, K., & Losardo, D. (2010). Twelve frequently asked questions about growth curve modeling. Journal of cognition and development, 11(2), 121–136. doi: https://doi.org/10.1080/15248371003699969
Gelman, A., & Rubin, D. B. (1992). Inference from iterative simulation using multiple sequences. Statistical science, 457–472. doi: https://doi.org/10.1214/ss/1177011136
Geweke, J. (1992). Evaluating the accuracy of sampling-based approaches to the calculations of posterior moments. Bayesian statistics, 4, 641–649.
Heidelberger, P., & Welch, P. D. (1983). Simulation run length control in the presence of an initial transient. Operations Research, 31(6), 1109–1144. doi: https://doi.org/10.1287/opre.31.6.1109
Lange, K. L., Little, R. J., & Taylor, J. M. (1989). Robust statistical modeling using the t distribution. Journal of the American Statistical Association, 84(408), 881–896. doi: https://doi.org/10.2307/2290063
McArdle, J. J., & Nesselroade, J. R. (2014). Longitudinal data analysis using structural equation models. American Psychological Association. doi: https://doi.org/10.1037/14440-000
Ntzoufras, I. (2011). Bayesian modeling using winbugs. John Wiley & Sons. doi: https://doi.org/10.1080/09332480.2012.685377
Raftery, A. E., Lewis, S., et al. (1992). How many iterations in the gibbs sampler. Bayesian statistics, 4(2), 763–773.
Tong, X., & Zhang, Z. (2017). Outlying observation diagnostics in growth curve modeling. Multivariate Behavioral Research, 52(6), 768–788. doi: https://doi.org/10.1080/00273171.2017.1374824
Tourangeau, K., Nord, C., Lê, T., Sorongon, A. G., Hagedorn, M. C., Daly, P., & Najarian, M. (2015). Early childhood longitudinal study, kindergarten class of 2010-11 (ecls-k: 2011). user’s manual for the ecls-k: 2011 kindergarten data file and electronic codebook, public version. nces 2015-074. National Center for Education Statistics.
Zhang, Z. (2016). Modeling error distributions of growth curve models through bayesian methods. Behavior research methods, 48, 427–444. doi: https://doi.org/10.3758/s13428-015-0589-9
Zhang, Z. (2021). A note on wishart and inverse wishart priors for covariance matrix. Journal of Behavioral Data Science, 1(2), 119–126. doi: https://doi.org/10.35566/jbds/v1n2/p2
Zhang, Z., Lai, K., Lu, Z., & Tong, X. (2013). Bayesian inference and application of robust growth curve models using student’s t distribution. Structural Equation Modeling: A Multidisciplinary Journal, 20(1), 47–78. doi: https://doi.org/10.1080/10705511.2013.742382
data=read.csv("example_data.csv",header = T) colnames(data)=c("ID",paste("math",rep(1:4),sep=’’)) data=data[,-1]
# The latent basis growth curve model model1 <- "model{ # specify the likelihood for (i in 1:nsubj) { for (j in 1:ntime) { # t error y[i, j] ˜ dt(mu[i, j], tauy, df) # normal # y[i, j] ˜ dnorm(mu[i, j],tauy) } } for (i in 1:nsubj){ mu[i,1] <- b[i,1] mu[i,2] <- b[i,1]+b[i,2] mu[i,3] <- b[i,1]+A3*b[i,2] mu[i,4] <- b[i,1]+A4*b[i,2] b[i,1:2] ˜ dmnorm(mub[1:2], taub[1:2,1:2]) } # specify the growth trajectory A3˜dnorm(0,1.0E-6) A4˜dnorm(0,1.0E-6) # specify priors mub[1]˜dnorm(0,1.0E-6) mub[2]˜dnorm(0,1.0E-6) taub[1:2, 1:2] ˜ dwish(Omega[1:2, 1:2], 2) sigma2b[1:2, 1:2] <- inverse(taub[1:2, 1:2]) tauy ˜ dgamma(0.001,0.001) sigma2y <- 1 / tauy df ˜ dunif(1,500) Omega[1,1] <- 1 Omega[2,2] <- 1 Omega[1,2] <- Omega[2,1] Omega[2,1] <- 0 } " # write model out writeLines(model1, "model1.txt") # set parameters, adaption, and MCMC chains parameters = c("mub","sigma2b","sigma2y","df", "A3","A4","dic")# Specify the estimated parameters adaptSteps =10000 # Adaptive period burnInSteps = 10000 # Burn-in period nChains = 2 nIter =40000 # The number of kept iterations nsubj = nrow(data) ntime = ncol(data) # create data set for JAGS model datalist = list(nsubj=nsubj,ntime=ntime,y=as.matrix(data)) # specify initial values inits <- list(list(mub=c(0.7,0.4), taub=structure(.Data = c(1,0,0,10),.Dim=c(2,2)), tauy=10,df=3), list(mub=c(0.7,0.5), taub=structure(.Data = c(2,0,0,8),.Dim=c(2,2)), tauy=15,df=5)) # run jags model set.seed(1234) out <- run.jags(model=model, monitor=parameters, data=datalist, n.chains=2, inits=inits, method="simple", adapt=adaptSteps, burnin = burnInSteps, sample=nIter, keep.jags.files=TRUE, tempdir=TRUE) # diagnostic geweke.diag(out$mcmc) # plots # trace plots and autocorrelation plots plot(out) # Summarize posterior distributions mcmcChain = as.matrix(out$mcmc) sum = summary(out$mcmc) # The linear LGCM model2 <- "model{ # likelihood for (i in 1:nsubj) { for (j in 1:ntime) { # t error y[i, j] ˜ dt(mu[i, j], tauy, df) } } # growth trajectory for (i in 1:nsubj){ mu[i,1] <- b[i,1] mu[i,2] <- b[i,1]+b[i,2] mu[i,3] <- b[i,1]+A3*b[i,2] mu[i,4] <- b[i,1]+A4*b[i,2] b[i,1:2] ˜ dmnorm(mub[1:2], taub[1:2,1:2]) } A3 <- 2 # linear change A4 <- 3 mub[1]˜dnorm(0,1.0E-6) mub[2]˜dnorm(0,1.0E-6) taub[1:2, 1:2] ˜ dwish(Omega[1:2, 1:2], 2) sigma2b[1:2, 1:2] <- inverse(taub[1:2, 1:2]) tauy ˜ dgamma(0.001,0.001) sigma2y <- 1 / tauy df ˜ dunif(1,500) Omega[1,1] <- 1 Omega[2,2] <- 1 Omega[1,2] <- Omega[2,1] Omega[2,1] <- 0 } " # Quadratic LGCM model3 <- "model{ # likelihood for (i in 1:nsubj) { for (j in 1:ntime) { # t error y[i, j] ˜ dt(mu[i, j], tauy, df) } } # growth trajectory for (i in 1:nsubj){ mu[i,1] <- b[i,1] mu[i,2] <- b[i,1]+b[i,2]+b[i,3] mu[i,3] <- b[i,1]+A3*b[i,2]+A5*b[i,3] mu[i,4] <- b[i,1]+A4*b[i,2]+A6*b[i,3] b[i,1:3] ˜ dmnorm(mub[1:3], taub[1:3,1:3]) } # linear change A3 <- 2 A4 <- 3 # quadratic change A5 <- 4 A6 <- 9 mub[1]˜dnorm(0,1.0E-6) mub[2]˜dnorm(0,1.0E-6) mub[3]˜dnorm(0,1.0E-6) taub[1:3,1:3] ˜ dwish(Omega[1:3, 1:3], 3) sigma2b[1:3, 1:3] <- inverse(taub[1:3,1:3]) tauy ˜ dgamma(0.001,0.001) sigma2y <- 1 / tauy df ˜ dunif(1,500) Omega[1,1] <- 1 Omega[2,2] <- 1 Omega[3,3] <- 1 Omega[1,2] <- Omega[2,1] Omega[1,3] <- Omega[3,1] Omega[2,3] <- Omega[3,2] Omega[2,1] <- 0 Omega[3,1] <- 0 Omega[3,2] <- 0 } "
# A linear LGCM model4 <- "model{ C <- 100000 lomega <- 0.5*loggam(3*(1+gamma)/2)-log(1+gamma) -3/2*loggam((1+gamma)/2) cgamma <- (exp(loggam(3*(1+gamma)/2)) /exp(loggam((1+gamma)/2)))ˆ(1/(1+gamma)) for (i in 1:nsubj) { for (j in 1:ntime) { # Exponential power zeros[i,j] ˜ dpois(zeros.mean[i,j]) zeros.mean[i,j] <- C-le[i,j] le[i,j] <- lomega-log(sqrt(sigma2y)) -cgamma*abs((y[i,j]-mu[i,j]) /sqrt(sigma2y))ˆ(2/(1+gamma)) } } # growth trajectory for (i in 1:nsubj){ mu[i,1] <- b[i,1] mu[i,2] <- b[i,1]+b[i,2] mu[i,3] <- b[i,1]+A3*b[i,2] mu[i,4] <- b[i,1]+A4*b[i,2] b[i,1:2] ˜ dmnorm(mub[1:2], taub[1:2,1:2]) } A3 <- 2 # linear change A4 <- 3 mub[1]˜dnorm(0,1.0E-6) mub[2]˜dnorm(0,1.0E-6) taub[1:2, 1:2] ˜ dwish(Omega[1:2, 1:2], 2) sigma2b[1:2, 1:2] <- inverse(taub[1:2, 1:2]) tauy ˜ dgamma(0.001,0.001) sigma2y <- 1 / tauy gamma ˜ dunif(-1,1) Omega[1,1] <- 1 Omega[2,2] <- 1 Omega[1,2] <- Omega[2,1] Omega[2,1] <- 0 } "
# A linear LGCM model5 <- "model{ C <- 100000 xi <- -sqrt(sigma2y)*(alpha/sqrt(1+alphaˆ2)) *sqrt(2/3.1415) for (i in 1:nsubj) { for (j in 1:ntime) { # Exponential power zeros[i,j] ˜ dpois(zeros.mean[i,j]) zeros.mean[i,j] <- C-le[i,j] e[i,j] <- y[i,j]-mu[i,j] # phi(): standard normal cdf # the log density of x is given by le[i,j] <- log(2)-log(sqrt(sigma2y)) +logdensity.norm((e[i,j]-xi)/sqrt(sigma2y),0,1) +log(phi(alpha*(e[i,j]-xi)/sqrt(sigma2y))) } } # growth trajectory for (i in 1:nsubj){ mu[i,1] <- b[i,1] mu[i,2] <- b[i,1]+b[i,2] mu[i,3] <- b[i,1]+A3*b[i,2] mu[i,4] <- b[i,1]+A4*b[i,2] b[i,1:2] ˜ dmnorm(mub[1:2], taub[1:2,1:2]) } A3 <- 2 # linear change A4 <- 3 mub[1]˜dnorm(0,1.0E-6) mub[2]˜dnorm(0,1.0E-6) taub[1:2, 1:2] ˜ dwish(Omega[1:2, 1:2], 2) sigma2b[1:2, 1:2] <- inverse(taub[1:2, 1:2]) tauy ˜ dgamma(0.001,0.001) sigma2y <- 1 / tauy alpha ˜ dunif(-5,5) Omega[1,1] <- 1 Omega[2,2] <- 1 Omega[1,2] <- Omega[2,1] Omega[2,1] <- 0 } "